-

Exploring the Potential of Viral Contributors to Alzheimer’s Disease
Alzheimer ‘s disease (AD) affects millions of Americans and causes significant morbidity and mortality. Although genetic determinants of AD have been a major focus of research over the last three decades, there is limited insight into co-factors that contribute to AD pathology and progression. Recent genetic associations implicating alterations in innate immunity to risk for AD, suggest that environmental factors, such as infection, may modulate brain immune function and could…
-

Integrating Big Data with DOCKET
The “first mile” problem of translational research is: how to integrate the multitude of dynamic small-to-large data sets that have been produced by the research and clinical communities, but that are in different locations, processed in different ways, and in a variety of formats that may not be mutually interoperable. Integrating these data sets requires significant manual work downloading, reformatting, parsing, indexing and analyzing each data set in turn. The…
-

Discovering and Annotating Transposable Elements
Repetitive DNA, especially that due to transposable elements (TEs), makes up a large fraction of most eukaryotic genomes. Over 58% of the human genome is derived from transposition and duplication of these prototypically selfish sequences. While only 1 in 20 humans have a new germline TE insertion, the somatic TE activity in tumors, during neural development in the brain, or during creation of pluripotent stem cells has a dramatic impact…
-

Sepsis Diagnostics
Each year, ~2,000,000 people in the U.S. develop sepsis and over 250,000 of these people die. Clinicians need advanced support when they make rapid, high-stakes decisions about sepsis for individual patients. Personalized risk stratification can help save lives and reduce unnecessary and potentially harmful care. Our lab, through a DoD funded study on post-operative sepsis, has developed blood biomarkers that can predict up to 3 days prior to diagnosis, with…
-
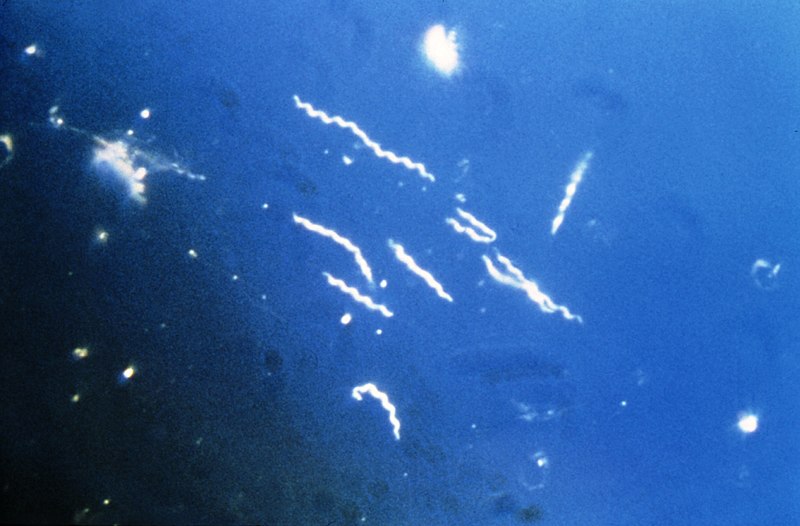
Lyme Disease Diagnostics
Lyme Disease Incidence has doubled over the past few decades with ~30,000 confirmed new case per year, which is likely an underestimate due to undiagnosed disease. Current diagnostic tests are only effective several weeks after infection since they rely on the host development of antibodies. Projects in our group are focusing on identifying biomarkers that can be diagnostic of infection at a much earlier time point. Furthermore, we are seeking…
-
Molecular Mechanisms of Influenza Infection
A major cause of morbidity/mortality during influenza virus infection is secondary bacterial co-infection. Indeed, bacterial co-infection is thought to have been the predominant cause of death during the 1918 influenza pandemic that killed over 50 million people. However, the molecular mechanisms underlying this synergy are not clearly defined. Our research has revealed that the mechanisms differ between influenza virus strains and include inhibition of lung epithelial repair processes and perturbations…
-
Patterns of Healthy Aging in the Gut Microbiome
Objectives: Characterize ‘healthy/unhealthy aging’ trajectories from the perspective of the gut microbiome across the adult human lifespan. Investigate changes in gut microbiome composition predictive of mortality and longevity in elderly populations. Study the reflection of the identified gut microbiome aging patterns in host physiology, primarily through blood analytes. Summarizing paragraph: Despite considerable progress in understanding the human gut microbiome, very little is known about how gut microbial changes across age…
-

Biomarkers for Post-Traumatic Stress Disorder
Blood based diagnostic marker for Post-traumatic stress disorder (PTSD) Post-traumatic stress disorder (PTSD) poses a significant burden on emotional and physical health and health care costs. According to US Department of Veterans Affairs, about 12% of Gulf war veterans, including those deployed to Iraq and Afghanistan, suffer from PTSD in a given year and about 30% of Vietnam veterans have had PTSD in their lifetime. The annual treatment cost for…
-

Extracellular Vesicles as a Diagnostic
Extracellular Vesicles in the Early Diagnosis of Small Cell Lung Cancer Lung cancer is the primary cause of cancer related deaths in the U.S and carries a 5-year survival rate of 17%. There are several reasons for these staggering facts including tumor heterogeneity, limited targeted therapeutics and late clinical presentation of disease. In the last several years, the development of therapeutics targeted to Epidermal Growth Factor Receptor (EGFR) and Anaplastic…
-
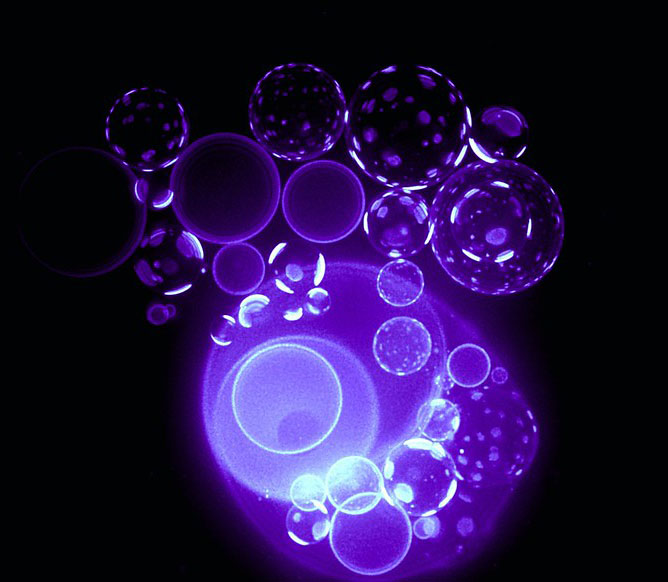

Analysis of Single Extracellular Vesicles
Microfluidics Array Based Sorting, Isolation, and RNA Analysis in Single Extracellular Vesicles Extracellular vesicles (EVs) such as microvesicles and exosomes are small membrane vesicles released by cells in the body. EVs are present in all biological fluids tested (e.g., blood, urine, cerebral spinal fluid) and contain various biomolecules including DNAs, RNAs, proteins and metabolites, and have been implicated as part of the cell-cell communication systems. Despite their importance, the current…
-
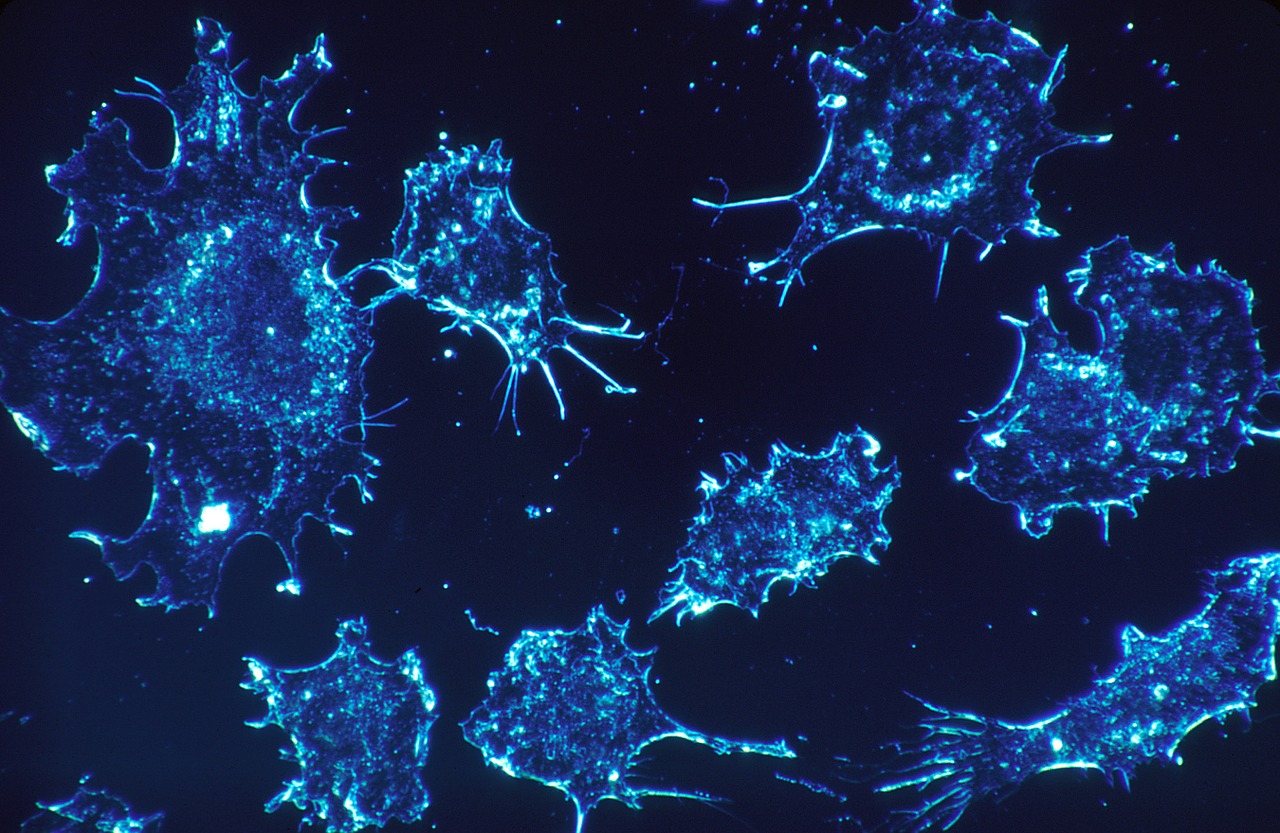
Predicting Cancer Phenotypes
This project aims to significantly advance our ability to study the complex mechanistic underpinnings of cancer using integrated multi-omic network models. Towards this goal, we are building metabolic networks relevant to cancer biology to serve as a mechanistic basis on which to ground statistical analyses of experimental multi-omic data. This collaborative systems-based approach for integrating and analyzing multi-omic information from regulatory network topology, signaling and metabolic pathways will lead to…
-

TReNa
The regulation of target genes by their transcription factors is complex and incompletely understood. Multiple signals involving core promoters, distal enhancers, epigenetic controls on chromatin accessibility, stochastic and cooperative binding on different times scales are all involved. Predictive modeling of these processes in fine detail and at scale is beyond our current capabilities. We know, however, that gene regulation usually includes two gross features which, independently, are poor predictors, but…
-
Personalizing Clinical Blood Tests
One of our focus areas is the personalized interpretation of clinical blood tests. We are exploring the extent to which deep-phenotyping data — especially genomics — can be used to personalize the interpretation of existing clinical laboratory measurements, ultimately improving their diagnostic accuracy. The integration of these personalized metrics with existing clinical decision-making offers the most straight-forward path to bring deep phenotyping into the clinic. In collaboration with physicians within…
-

Manifestation of Polygenic Risk Scores
Transitions from health to disease are characterized by dysregulation of biological networks under the influence of genetic and environmental factors, often over the course of years to decades before clinical symptoms appear. Understanding these dynamics has important implications for preventive medicine. However, progress has been hindered both by the difficulty of identifying individuals who will eventually go on to develop a particular disease, and by the inaccessibility of most disease-relevant…
-

PHEWAS Approach to Study Alzheimer’s Disease
Genetics play an important role in late-onset Alzheimer’s Disease (AD) etiology. As AD develops over decades, elucidating the biological effects of AD-associated genetic variation across the adult lifespan may illuminate underlying processes in the causal pathway for AD, and potentially provide novel mid-life targets for behavioral and drug intervention. To address this, we are utilizing high dimensional, high-quality measurements from thousands of participants in a wellness program, aged 18 to…
-

PARreto Task Inference Analysis on Multi-Omic Data
Objectives: Analyzing multi-omic data using Pareto task inference method Characterize archetypes of health states and reveal the trade-offs in the human wellness space Find biomarkers for early detection of transitions from health to disease state It is a challenge to answer questions like: Why some people develop a disease, react to a specific treatment and/or develop severe side-effects while others don’t. In order to explain these occurrences, one has to…
-
Transcriptional Regulatory Networks in Alzheimer’s Disease
We are using Transcriptional Regulatory Networks (TRNs) to better understand the causes and progression of Alzheimer’s disease. Our approach starts with measurements of gene expression in the brain, from which we create models that help to understand the changes in transcriptional regulatory structure that occur in Alzheimer’s disease. We are developing methods that help to identify cell-type specific changes in gene expression, as well as changes in gene regulation by transcription factors….
-
Predicting Gut Microbiome Diversity from Blood
Blood metabolome predicts gut microbiome α-diversity in humans Objectives: Study the reflection of gut microbiome in the bloodstream Find blood biomarkers for low, potentially problematic, gut microbiome diversity Study the complex interconnections between host metabolism, gut microbial composition, and human health. Summarizing paragraph: The microbes in our gut are important to our health, but we do not yet know how to tell if a particular mix of microbes is “healthy”….
-

Genotype, Genome and Data Fingerprints
Genotype, Genome and Data Fingerprints Soon, millions of individual human genomes with rich phenotype data will be available for analysis, posing a data management challenge and offering significant discovery opportunities. Rich genomic and phenomic knowledge will help improve our understanding of genome structure, function and evolution, and will translate into actionable opportunities for improving health and wellness. We have developed several algorithms and methods for studying and visualizing personal genome…
-

Biological Age
Biological Age, derived from molecular and physiological measurements, has been proposed to better predict mortality and disease than chronological age. We have leveraged longitudinal deep phenotyping data from a real world cohort to develop a computed estimate of biological age. In our studies, biological age relative to chronological age, was elevated in the presence of chronic disease, providing evidence for the value of our approach. Since biological age is modifiable,…
Privacy Overview
This website uses cookies so that we can provide you with the best user experience possible. Cookie information is stored in your browser and performs functions such as recognising you when you return to our website and helping our team to understand which sections of the website you find most interesting and useful.



